mechanism of expanding bacteria revealed
Mechanism of expanding bacteria revealed
Published on: 15 June 2022
A new study published in Nature has identified a potential Achilles heel in the protective layers surrounding Gram-negative bacteria that could aid in the development of next-generation antibiotics.
The study, carried out jointly by Professor Waldemar Vollmer and Dr Federico Coronaat Newcastle University, alongside Professor Colin Kleanthous and Dr Gideon Mamouin the Department of Biochemistry at the University of Oxford, shows that Gram-negative bacteria depend on the cell wall to synchronise building of the outer membrane.
The World Health Organization (WHO) has declared Antimicrobial Resistance (AMR) one of the top 10 global public health threats. Some bacteria have already become resistant to all known antibiotics. Particularly problematic are multidrug resistant Gram-negative bacteria such as Escherichia coli, Pseudomonas aeruginosa and Klebsiella pneumoniae that cause pneumonias and sepsis. Their outer membrane resides beyond the cell wall and excludes many classes of antibiotics that would otherwise target it.
The research reveals that the cell wall, which is composed of a tough material known as peptidoglycan, exerts surprising control over where new proteins are introduced into the outer membrane by an essential biogenesis protein known as BamA. This helps bacteria coordinate these layers, which is crucial for the way they grow.
Professor Vollmer from the Centre for Bacterial Cell Biology at Newcastle University Biosciences Institute explains: “Bacteria are tiny yet have a very high internal pressure, like a car tyre. They have a strong cell wall to withstand this pressure and prevent them bursting while still allowing them to grow and divide. What we have revealed for the first time is as they grow, how the process of wall expansion is linked to that of the outer membrane beyond it.”
The research revealed that the age of the mesh-like peptidoglycan – composed of amino acids and sugars – is the key factor. ‘Old’ peptidoglycan that occurs primarily at the poles (ends) of cells completely shuts down new outer membrane protein biogenesis. By contrast, ‘new’ peptidoglycan that occurs primarily at sites where cells are about to divide, allows the greatest amount of outer membrane growth. This simple yet elegant mechanism ensures bacteria maintain a tight control over both cell envelope layers.
Professor Kleanthous from Oxford added: “We never suspected that Gram-negative bacteria were so reliant on the cell wall to coordinate growth of the outer membrane. Disrupting this cross-talk would literally ‘open up’ Gram-negative bacteria, making them vulnerable to antibiotics that the outer membrane otherwise excludes.”
The work was supported by the European Union’s Horizon 2020 programme, the European Research Council, the Wellcome Trust, the Biotechnology and Biological Sciences Research Council and the Medical Research Council.
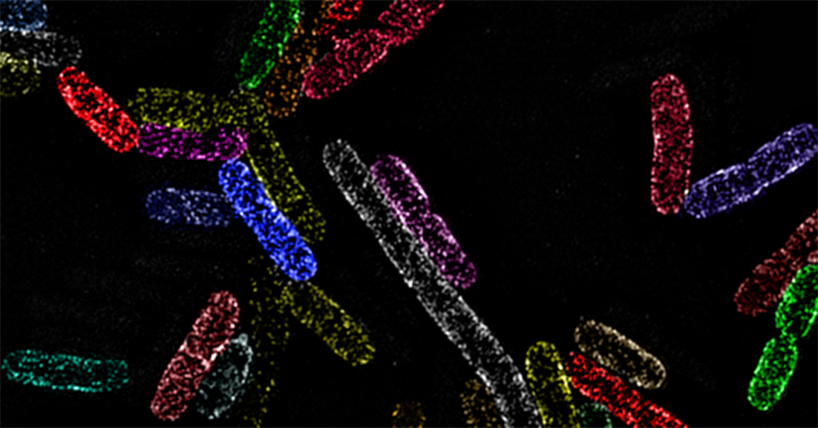
Figure caption and publication
Figure Caption. Image shows clusters of the essential protein BamA in the outer membrane of the bacterium Escherichia coli. The new study by Mamou and colleagues demonstrates that BamA’s ability to introduce new proteins into the membrane, which are needed for growth, is strictly controlled by the cell wall that sits immediately beneath it. Image courtesy of Dr Gideon Mamou, University of Oxford.
Publication: Peptidoglycan maturation controls outer membrane protein assembly. DOI: 10.1038/s41586-022-04834-7. Nature.
Latest News
Heading 3 example
Text only. For subheading use ‘Heading 3’
Lorem ipsum dolor sit amet, consectetur adipiscing elit. Donec vel sapien lectus. Aliquam consectetur vitae tortor mattis sodales. Donec id quam id nulla tristique vestibulum. Ut vitae orci aliquam massa varius dapibus. Vivamus aliquet, lorem sit amet semper ultricies, ex tortor molestie felis, at ultrices tellus ligula vitae est. Sed gravida tortor sapien, in iaculis quam vestibulum vel. Duis et quam nec metus pharetra placerat. Donec in tellus pretium ex sagittis posuere. Nunc varius, libero at suscipit commodo, magna lacus facilisis velit, sed pellentesque magna eros vel dolor. Donec pretium neque ultrices, condimentum turpis non, porttitor neque. Suspendisse fermentum at lectus scelerisque mattis. Nam augue justo, iaculis quis euismod et, viverra in elit. Aliquam vitae justo malesuada, sodales urna et, aliquam nunc. Nulla placerat neque quis odio molestie, mattis bibendum turpis pellentesque.



